Metadata: The 'How' and 'Why' Behind Your Science
This is the first post in a series that will cover what we’ve learned since starting Kaleidoscope: major challenges in the field of bio R&D, opportunities for accelerating science, and the decisions/philosophies underpinning our work. If anything in the posts resonates with you, please reach out!
Summary points
- Metadata is a way to preserve key artifacts about all the data and knowledge you generate. It is critical for understanding the context of scientific projects — metadata provides information about what experiments or tests you ran, how you ran them, and why.
- Good data hygiene and metadata capture has compounding gains — it allows you to systematically build on past knowledge and more effectively scale your science.
- Kaleidoscope gives teams a simple and intuitive framework to capture metadata in the same place they plan and collaborate on their science.
Over the last 12 months, we connected with hundreds of researchers, engineers, and scientific teams working in bio, to try and understand the problems they are facing. We heard about challenges with data analysis, technical barriers to properly leveraging cloud compute, pains with scaling science, and numerous other topics. So why, then, did we start by bringing metadata to the forefront of our product at Kaleidoscope? And, more broadly, what is metadata and why should you care?
A common thread across all of the conversations we had was that keeping track of a growing body of experiments is exceptionally difficult. Not only are experiments themselves complex projects, but the volume of output and the rate of sharing (internal and external) are exploding. These (and other) factors mean that the challenges of tracking work are only compounding over time. Important experimental information is lost along the way, making it really hard to reproduce past results or generate novel insights in a reliable and scalable way. Enter: metadata.
Metadata is the context of experiments and scientific outputs — it allows you to make sense of the data you accrue over time. What protocols were used, what data was generated (and who interacted with it), how experiments were related to each other, what conclusions you reached, and much more.
Capturing metadata allows you to begin to systematically learn about your science as a whole, rather than viewing it as a collection of fragmented and disjointed pieces.
From here, it becomes possible to start using past learnings to generate novel predictions and insights that may not be visible to the human eye; at scale, this is a powerful thing. In fact, metadata is a key ingredient to process optimization or AI/ML implementation, which is something bio orgs are increasingly able and looking to do.
Effectively tracking and leveraging metadata is not easy — even ML-heavy teams (outside of biotech) struggle with this. Training and tracking models and their performance against different test sets is often much less systematic than it should be; model parameters are hard-coded instead of separately tracked, model lineage is not properly recorded, and so on. When you layer on the fact that early R&D is, by definition, chaotic and often unstructured, things quickly become messy and data hygiene can slip into an afterthought. Some would even argue that this chaos and freedom to operate in an unconstrained way is necessary for breakthrough science to happen (and they’re not entirely wrong).
At Kaleidoscope, we made metadata central in the product to encourage best practices from the field of data science, while allowing for the natural unpredictability of R&D.
We deeply believe that being systematic and thoughtful about what you’re tracking (and why) should be core to how you do science, and not an afterthought.
And, despite the challenges mentioned above, we’re ardent believers that good data hygiene and unconstrained scientific creativity can coexist, given the right product. For us, this means:
- Giving people simple and intuitive tools
- Making it easy to go from ad-hoc to structured and templated work
- Creating an environment where metadata capture sits right next to the science
- Encouraging integrations across other software in the stack and minimizing repeat work
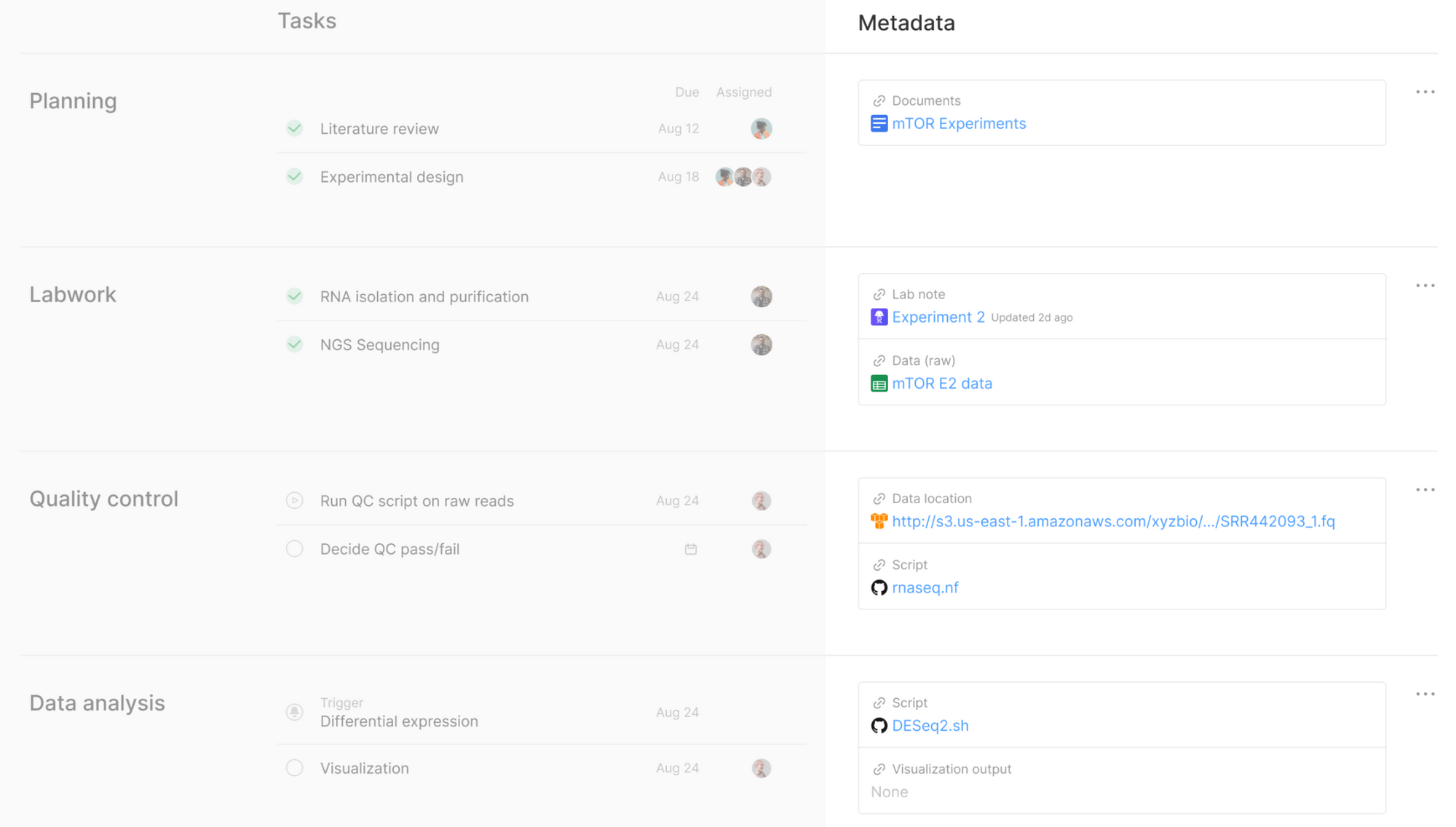
With Kaleidoscope, we prompt people to think about the context of their work and give them a way to consistently capture metadata. Tracking metadata in Kaleidoscope also enables visibility across teams and tools: context and key artifacts can be pulled from wherever they sit, hand-offs between team members become easier, and different functional teams (eg wet lab and dry lab) can align more seamlessly. In turn, this becomes a powerful utility underpinning a whole range of exciting, downstream features, such as data health checks or scientific versioning (more on that later). And, over time, we can introduce ways to automate how this metadata is captured (and analyzed), so that you don’t have to spend much time doing so.
If you’re still not convinced of the importance of metadata today, consider the fact that the benefits of capturing metadata are not just ‘limited’ to future-looking, breakthrough-science use-cases. Let’s say a new team member joins or someone changes teams: how do they quickly and effectively catch up with the work their team/org has done to date? How do you give people a way to easily connect the dots between what was done and why, so that they have context to the work they are doing and can more quickly contribute to the science and discovery process? Metadata tagged to projects is a way for people to have a map of the science they are working on.
With all that said, effectively implementing metadata capture, internally championing its importance, and properly leveraging the results is hard. R&D can be messy. Humans have limited bandwidth. And the value of processes that pay long-term dividends can be hard to quantify in the present. At Kaleidoscope, we’re working hard to drastically simplify this and give you tools that make it painless.
If you want to chat more about anything we wrote, or you’re interested in finding a way to work together, let us know!
